Liquid Biopsies to Detect Cancer Could Boost Annual Sequencing Volume By 40-Fold
Next-generation DNA sequencing (“NGS”) is creating a new wave of diagnostic tests for cancer. The level of detail offered by NGS tools is helping researchers make important connections between the biology of tumors and the successful treatment of cancer.
Cancer is a dynamic and elusive disease. Its genetic profile mutates over time, sometimes in response to therapy. Consequently, tumors have to be monitored regularly for molecular changes. While tissue biopsies have been the standard of care historically, they are invasive and often not successful or reliable because of a tumor’s location or metastasis. Meanwhile, sequencing costs have dropped to a point that clinicians now can track the evolution of advanced tumor tissues noninvasively over time by sampling and testing blood: a liquid biopsy.
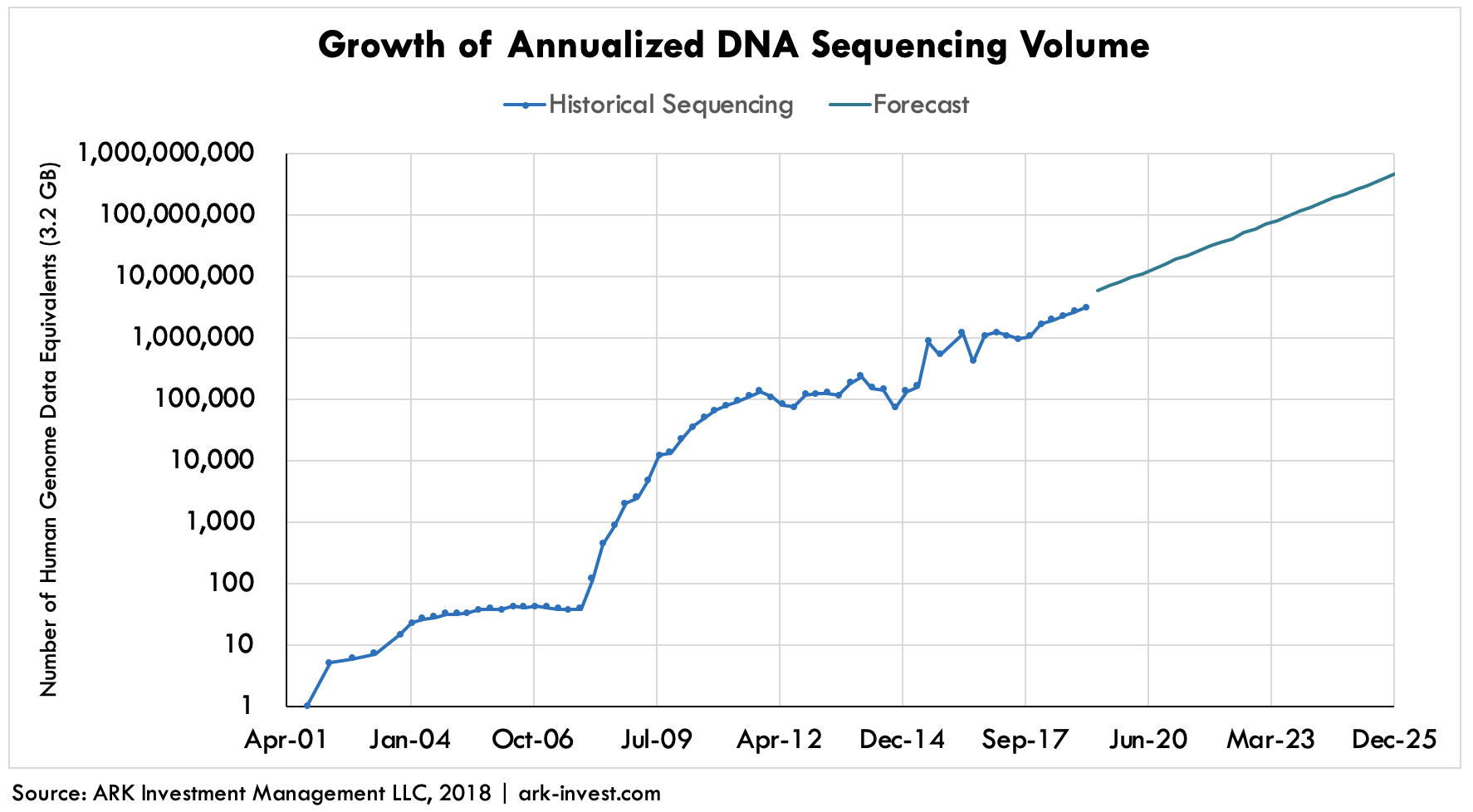
With liquid biopsies (“LBs”), clinicians are searching for trace amounts of tumor DNA (“ctDNA”) circulating in the bloodstream that, when sequenced, can guide treatment decisions. While still in development, liquid biopsies will be tuned not only to detect cancer in its earliest stages but also to monitor patients in remission. Cancer is easier to treat if discovered early, but the sequencing required for early stage tumors is more intensive, and more cost prohibitive, than that in later stages. Recurrence rates for most forms of cancer are harrowing, and for some tumors like gliomas inevitable. According to our estimates, if only the patients at risk of recurrent cancer were to be sequenced on an annual basis, the volume of genetic data produced would leap by 40-fold—an amount equivalent to that expected when 100 million whole human genomes will be sequenced globally in 2023, as shown below.
Screening the body for cancer involves a targeted form of NGS that focuses on a subset of cancer-causing genes, also known as oncogenes, that can mutate and cause tumors. Currently, more than 6 million oncogenic mutations span a region equivalent to just 2% of the human genome. Companies that have developed liquid biopsies for single cancer indications, such as Guardant Health (GH), focus on regions even smaller. Guardant’s liquid biopsy test for advanced non-small cell lung cancer, for example, focuses on mutations comprising 78,000 base pairs of DNA—corresponding to only 0.0025% of the human genome. Although these target regions are small, clinicians sequence the DNA so many times that the data generation can exceed that associated with one whole human genome.
Tumor cells shed ctDNA into the bloodstream in extraordinarily small quantities. Most NGS diagnostics split the DNA into small fragments and sequence it many times to reduce the probability of random errors. This multiple, or sequence coverage, can vary widely. In cancer patients, the ratio of ctDNA to healthy DNA can drop below one in one hundred. To detect this trace amount of ctDNA in the bloodstream, the tumor’s signal must be amplified. GH reports a liquid biopsy coverage value of 8,000X for lung cancer, meaning that the genetic target is read 8,000 times prior to analysis. In pan-cancer liquid biopsies, Grail has reported coverage of 60,000X. Consequently, one liquid biopsy for monitoring pan-cancer recurrence can have a sequencing intensity equivalent to mapping 40 whole human genomes.
Just to monitor the recurrence of cancer, liquid biopsies could be the reason that NGS volumes will scale from 2.4 million in 2018 to 100 million genome-equivalents per year, pushing costs down towards the $100 per genome predicted by Wright’s Law, by 2023.

 Actively Managed Equity
Actively Managed Equity Overview: All Strategies
Overview: All Strategies Investor Resources
Investor Resources Indexed Equity
Indexed Equity Private Equity
Private Equity Digital Assets
Digital Assets Invest In The Future Today
Invest In The Future Today
 Take Advantage Of Market Inefficiencies
Take Advantage Of Market Inefficiencies
 Make The World A Better Place
Make The World A Better Place
 Articles
Articles Podcasts
Podcasts White Papers
White Papers Newsletters
Newsletters Videos
Videos Big Ideas 2024
Big Ideas 2024


